Structural Biology
The Structural Biology Research group (Professor Trevor Greenhough, Dr Annette Shrive) aims to characterise the molecular mechanisms by which molecules of the innate immune system recognise and bind to their natural targets and effect clearance through interaction with components of immune system pathways.

This structural immunology includes ligand/pathogen recognition by the pentraxins CRP and SAP from human and a variety of other sources including Limulus, rat and fish; by collectins including hSP-D and hSP-A and by other proteins targeted by our collaborators for study. Of particular interest in the group is the pathogen-host interaction, with a particular emphasis on humans and mammals but also extending through to fish and to plants. In addition to crystallography, our structural investigations utilise complementary techniques, both in-house and with collaborators, including SRCD, cryoEM, SAXS, carbohydrate-type ligand manipulation and analysis.
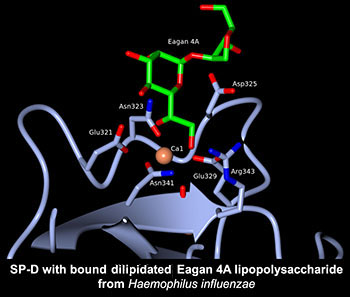
Human lung surfactant protein D (hSP-D) can directly interact with carbohydrate residues on pulmonary pathogens and allergens, stimulate immune cells, and manipulate cytokine and chemokine profiles during the immune response in lungs. The high resolution crystal structures, both native and ligand bound, of a therapeutically active recombinant fragment of SP-D define the fine detail of the mode and nature of carbohydrate recognition and provide insights into how a small fragment of human SP-D can bind to whole pathogens and at the same time recruit and engage effector cells and molecules of humoral immunity. Current emphasis is continued structural investigation of microbial recognition by hSP-D and by the major lung surfactant protein hSP-A, provided by our UCL collaborators, in order to characterise recognition properties and the pathogen neutralisation synergy between hSP-A and hSP-D.
Key references:
- Williams et al. (2026) J. Biol. Chem. 302(4), 111307.
- Paterson et al. (2019) Journal of Biological Chemistry, 294, 17155-17165.
- Littlejohn et al. (2018) PLoS One. 18;13(6):e0199175.
- Clark et al. (2016) Infection and Immunity. 84, 1585-1592.
- Shrive et al. (2009) Journal of Molecular Biology. 394, 776-788.
- Kishore et al. (2004) Immunology Letters 95, 113-128.
- Kishore et al. (2004) Trends in Immunology 25, 551-561.
- Shrive et al. (2003) Journal of Molecular Biology. 331, 509-523.
Human C-reactive protein (CRP) is a trace plasma protein which exhibits rapid increases in concentration of up to 1000-fold in response to tissue damage and inflammation. As the classical acute-phase reactant, CRP is used almost universally as a clinical indicator of inflammation and underlying infection. The host defence functions of human CRP depend to a great extent on its ability to activate the classical complement pathway and a major current interest is in structural insight into its ligand recognition and effector function using a variety of complementary techniques (x-ray crystallography, cryo-EM, SAXS) in collaboration with colleagues at Diamond Light Source and Midlands Regional Cryo-EM Facility.
Key references:
- Williams et al. (2020) Frontiers in Immunology, 11, 115.
- Shrive et al. (1996) Nature Struct. Biol. 3, 346-354.
- Agrawal et al. (2001) J. Immunol. 166, 3998-4004.
- Ramadan et al. (2002) Acta Cryst. D58, 992-1001.
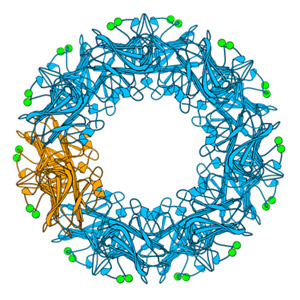 Our work on pentraxins has included proteins from various species. Work on the serum-amyloid-P-component-like pentraxin from Limulus polyphemus includes determination of sequence and three-dimensional structure and complementary studies in collaboration with University of California. Our work on C-reactive protein (CRP) in carp, with Prof Dave Hoole, was part of two successive EU Marie Curie training networks with European partners, that sought to establish the role of CRP in fish and its potential use as a biomarker of health in a similar manner to the use of CRP in humans in the clinical setting.
Our work on pentraxins has included proteins from various species. Work on the serum-amyloid-P-component-like pentraxin from Limulus polyphemus includes determination of sequence and three-dimensional structure and complementary studies in collaboration with University of California. Our work on C-reactive protein (CRP) in carp, with Prof Dave Hoole, was part of two successive EU Marie Curie training networks with European partners, that sought to establish the role of CRP in fish and its potential use as a biomarker of health in a similar manner to the use of CRP in humans in the clinical setting.
Key references:
- Pionnier et al. (2014) Fish and Shellfish Immunology 39, 285-295.
- Pionnier et al. (2013) Fish and Shellfish Immunology 34, 819-831.
- Shrive et al. (2009) Mol. Biol.386, 1240-1254.
- MacCarthy et al. (2008) Developmental and Comparative Immunology 32, 1281-1289.
- Cartwright et al. (2004) Developmental and Comparative Immunology28, 113-125.
- Tharia et al. (2002) Mol. Biol.316, 583-597.
- Shrive et al. (1999) J. Mol. Biol. 290, 997-1008
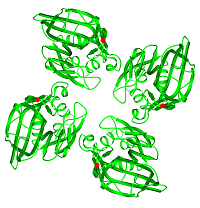
FIBCD1 (a collaborative project with the University of Southern Denmark) is a tetrameric plasma membrane protein that uses a fibrinogen-like recognition domain (FReD) for pattern recognition of acetyl groups on chitin. The x-ray structures of the FIBCD1 FReD reveal how FIBCD1 binds acetylated pathogen associated molecular patterns (PAMPS) and endogenous glycosaminoglycans, combining versatility with conservation to recognise its targets.
Key reference:
- Williams et al. (2024) J. Biol. Chem. 300(1), 105552
- Shrive et al. (2014) J. Biol. Chem. 289, 2880-2887.



